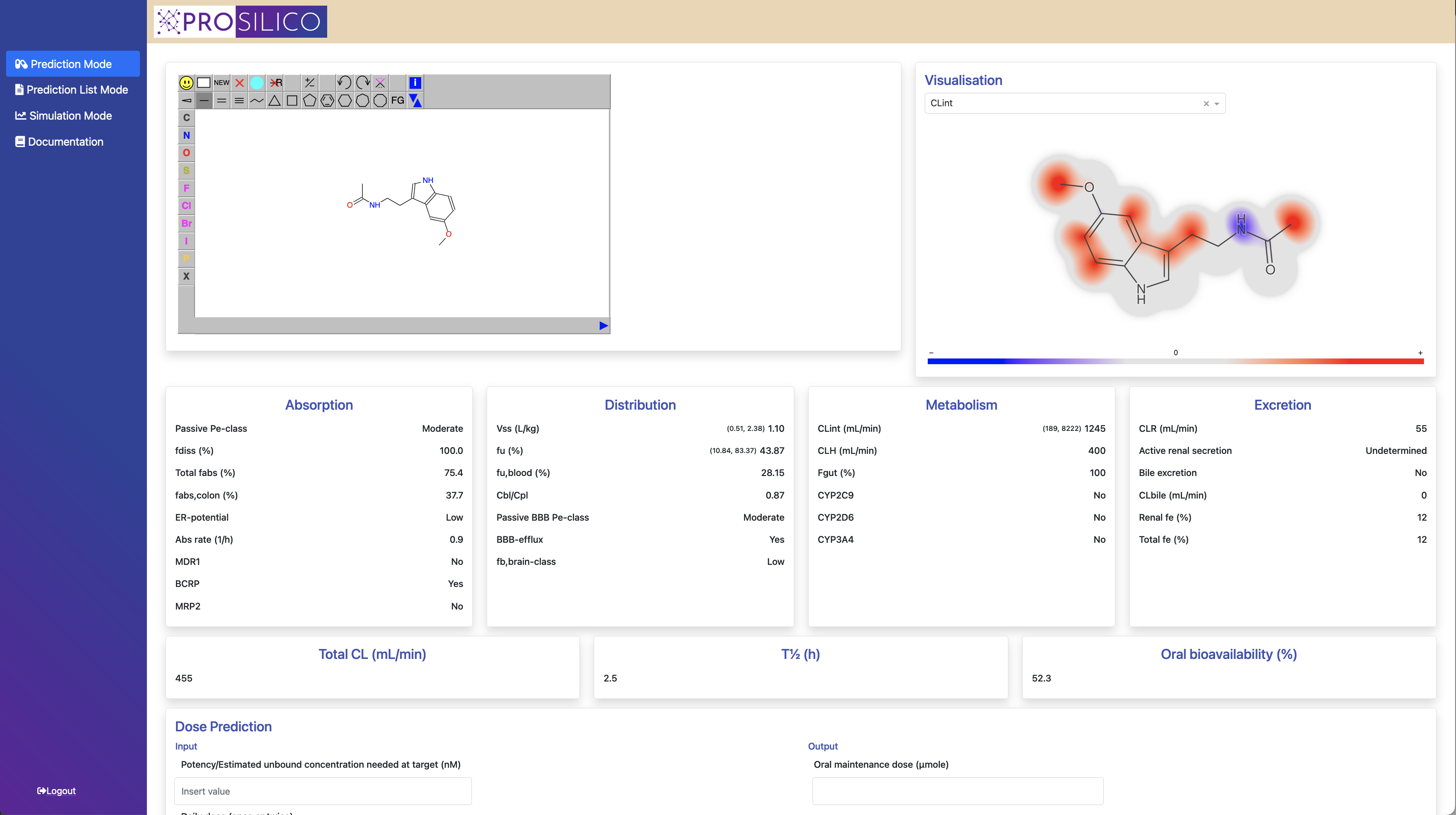
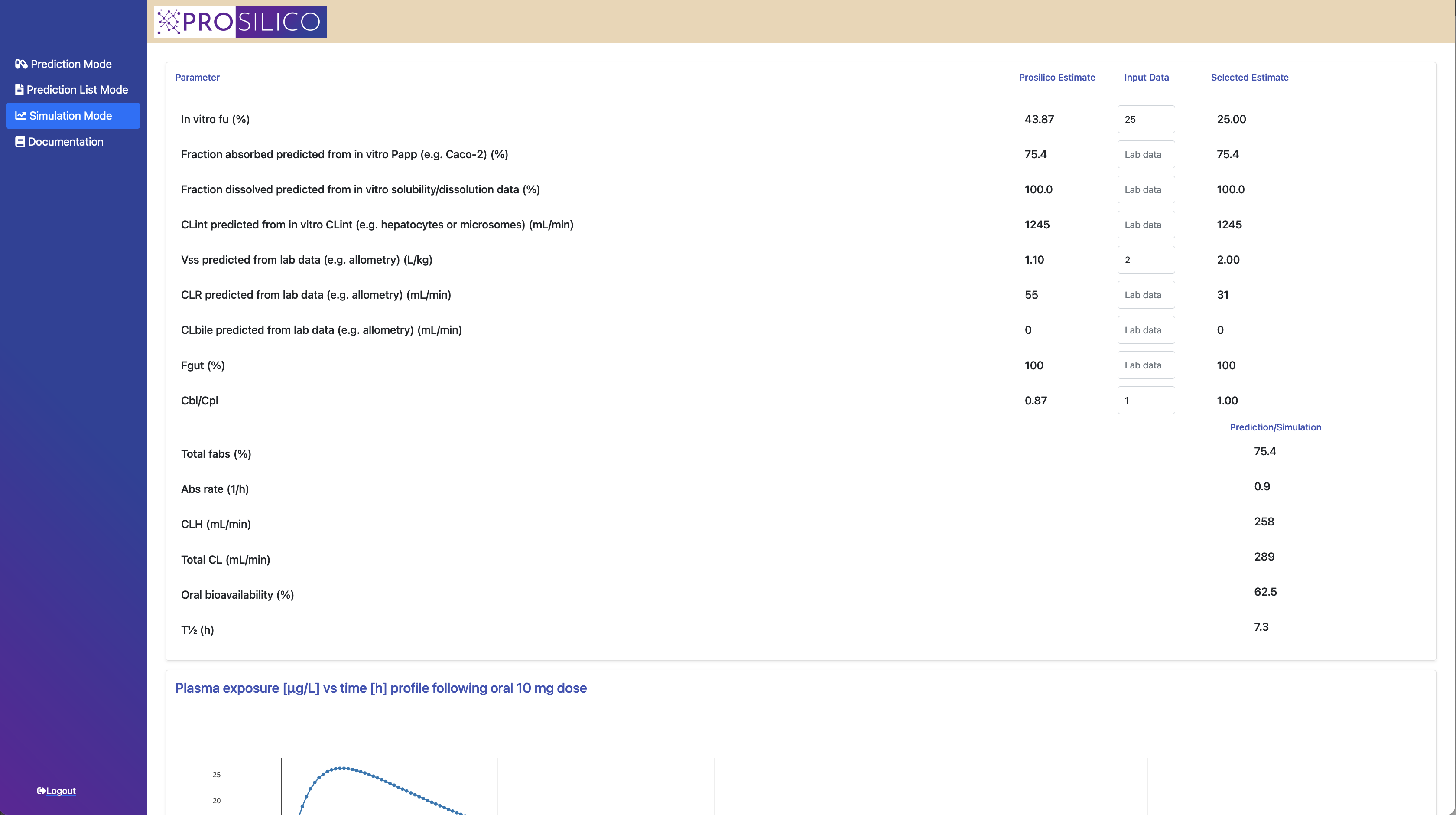
With the extensively validated ANDROMEDA by Prosilico software you will be able to predict, simulate and optimize the human clinical ADME/PK, exposures and doses of drug candidates, metabolites and other types of chemicals.
You get 30 ADME/PK parameters at your fingertips (see figure below), and performance and throughput superior to that of laboratory methods. You can predict batches of compounds and also include your own laboratory data.
The software is applicable for modern drugs, which often have more complex/multi-mechanistical characteristics, and works for compounds with low solubility and recovery, low CLint, significant excretion, gut-wall extraction and extensive efflux.